

lu·te·fisk \´lüd·e¸fisk, ´
lüe-\ or lut·fisk \´lüt¸f-\ also
lu-
de·fisk \´lüde-\ or lud·fisk \´
lüd¸f-\ n -s [lutefisk fr. Norw,
fr. lute
to wash in lye solution + fisk fish; lutfisk fr. Sw, fr.
luta
to wash in lye solution + fisk fish; ludefisk & ludfisk fr.
Dan
ludfisk fr. lude to wash in lye solution + fisk fish; Norw
lute, Sw luta, Dan lude akin to ON laug bath, hot spring;
Norw
& Sw & Dan fisk fr. ON fiskr fish -- more at LYE, FISH]
: stockfish that has been soaked in lye water, skinned,
boned,
and boiled
|
Lutefisk is software for the de novo interpretation of peptide CID spectra
High quality tandem mass spectra of peptides are often obtained for which no exact database match
can be made. Consequently, we are faced with the question of whether the protein under
investigation is novel, or if the non-matching spectra are due to less exciting prospects such
as inter-species variation, database sequence errors, or unexpected proteolytic cleavages.
To begin addressing this problem we perform a de novointerpretation of the CID spectra
using the computer program Lutefisk; however, any such interpretations nearly always yield
multiple sequence candidates, where it is often difficult or impossible to distinguish the
correct sequence from the incorrect ones. The variations between candidate sequences are
often minor and typically involve dipeptide inversions, swapping of dipeptides of the same
mass, replacements of dipeptides with single amino acids of the same mass, and replacements
of amino acids by dipeptides of the same mass. We use the multiple sequence candidates produced
by Lutefisk as query sequences in a second program, CIDentify. CIDentify is a version of
Bill Pearson's FASTA algorithm modified by Alex Taylor to accommodate MS nuances such as
multiple query sequences, ambiguous dipeptides and isobaric mass equivalencies.
Download:
Lutefisk has now been released under the GNU General Public License (GPL). ANSI C Source code which can be
compiled for MacOS X, Windoze, Linux, Solaris, Irix, AIX, or OSF and a Windoze binary can
be downloaded from sourceforge.
Source code and executables for CIDentify can be obtained from the CIDentify folder at the
FASTA site at the University of Virginia.
User's Guide:
References:
- R. S. Johnson and J. A. Taylor (2002) "Searching sequence databases via de novo peptide sequencing by
tandem mass spectrometry", Mol. Biotechnology Vol. 22, No. 3. (November 2002), pp. 301-315.
- J. A. Taylor and R. S. Johnson (2000) "Implementation and uses of automated de novo peptide sequencing
by tandem mass spectrometry". Anal. Chem. 73:2594-2604. [PDF (login required)]
- R. S. Johnson and J. A. Taylor (2000) "Searching sequence databases via de novo peptide sequencing by
tandem mass spectrometry", "Protein and Peptide Analysis: Advances
in the Use of Mass Spectrometry", 146:41-61 (Methods in Molecular Biology series, Humana Press).
- J. A. Taylor and R. S. Johnson (1997) "Sequence database searches via de novo peptide
sequencing by tandem mass spectrometry". Rapid Comm. Mass Spec.. 11:1067-1075. [PDF (account required)]
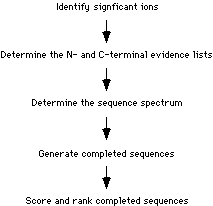
How Does It Work?:
This is a general flowchart depicting the major portions of the algorithm:
Follow this link for an
illustrated example
of how the program works.
License / Contact Info:
This program is free software; you can redistribute it and/or
modify it under the terms of the GNU General Public License
as published by the Free Software Foundation; either version 2
of the License, or (at your option) any later version.
This program is distributed in the hope that it will be useful,
but WITHOUT ANY WARRANTY; without even the implied warranty of
MERCHANTABILITY or FITNESS FOR A PARTICULAR PURPOSE. See the
GNU General Public License for more details.
You should have received a copy of the GNU General Public License
along with this program; if not, write to the Free Software
Foundation, Inc., 51 Franklin Street, Fifth Floor, Boston, MA 02110-1301, USA.
Questions? Problems?
Other tidbits:
You can find Rich's ABRF MS/MS Interpretation Tutorial
here.
Need a lutefisk
recipe?
Mass Spectrometry Poetry / Haiku Corner
b and y ions
low energy CID
mystery sequence
 -RSJ -RSJ
|
|
instrument makers
don't be anal retentive
i need file structures
 -RSJ -RSJ
|
|
valleys with no peaks
the smooth and tranquil landscape
of dirty samples
 -Ioannis Papayannopoulos
-Ioannis Papayannopoulos
|
|
your database search
has yielded some hits; alas
HGS has patents
 -Ioannis Papayannopoulos
-Ioannis Papayannopoulos
|
|
peptide fragments
many ions, much confusion
trees in a forest
 -Ioannis Papayannopoulos
-Ioannis Papayannopoulos
|
|
beauty is skin deep
but samples with keratin
are just plain ugly
 -J. Alex Taylor
-J. Alex Taylor
|
|
maldi-tof spectra
so many peptide masses
what is all this crap
 -J. S. Richar
-J. S. Richar
|
|
like matthias mann
we wish to become famous
but we never will
 -J. S. Richar
-J. S. Richar
|
|
Lutefisk Home Page
 Sherpa Home Page
Sherpa Home Page
